IMAGIC
Description | Working with IMAGIC | Manuals | Price List | Download | IMAGIC FormatsDescription
|
IMAGIC is a high end environment for the analysis of images,
spectra and other multi-dimensional data-sets.
IMAGIC can also be used in light and raster-tunnelling microscopes, computer tomographs, FT-IR spectrometers and other signal collecting devices. Even protein sequences from full data banks have been processed. |
|
|
Multi-dimensional mixed-radix Fourier transforms (limited by disk
space only),
correlation analysis,
alignments,
three-dimensional reconstruction from projections
and
morphometry
are all standard procedures in this floating point oriented data processing system.
The multivariate statistical pattern recognition procedures (MSA) of IMAGIC are geared at extracting the essential information from large and complex data-sets in various disciplines like biology, medicine and environmental sciences. These MSA procedures render huge, noisy data sets manageable and understandable. A friendly and transparent user-interface with local interactive help at all levels completes the application-oriented user extendible IMAGIC. |
|
The three-dimensional data processing and angular reconstitution modules allow the three-dimensional reconstruction of biological macromolecules with arbitrary point-group symmetry from their two-dimensional electron microscopical projections. By exploiting the different orientations of the molecules in the embedding medium, one may extract 3D information from the data without ever collecting tilt series in the microscope. The simplicity of this approach allows fast and unproblematic 3D analysis of biological macromolecules with high resolution.
In the latest IMAGIC-4D version all programs will automatically loop over stacks of 2D images or 3D volumes present in an IMAGIC input file. All operations are usually in-core ensuring fast processing avoiding unnecessary input/output operations.
All time-critical programs exploit MPI parallelization allowing for millions of input images to be processed efficiently.
IMAGIC is running on various Linux / UNIX computers as well as under MS Windows and Intel based Mac OS X computers.
IMAGIC Modules

| BASIC: Modules for standard image processing and analysis | ||||||||||||||||||||||||

| EM: Modules for electron microscopy (CTF correction, particle picking,alignments...) | ||||||||||||||||||||||||

| MSA: Module for multivariate statistical analysis and classification | ||||||||||||||||||||||||

| THREED/ANGREC: Modules for angular reconstitution and 3D/4D image analysis and reconstruction | ||||||||||||||||||||||||

| MORFO: Modules for morphometrical calculations | ||||||||||||||||||||||||

| SPECTRA: Module for 1-D/spectra analysis and image processing |

|
Terminal mode:
IMAGIC is running in a line-by-line mode in terminal/command window. 
Give IMAGIC in a terminal window to start IMAGIC in this mode. |
||||||||||||||||||||||||

|
guiIMAGIC:
You can run each IMAGIC command in GUI oriented mode. Various buttons allow for browsing files, context sensitive help, display, plot etc. 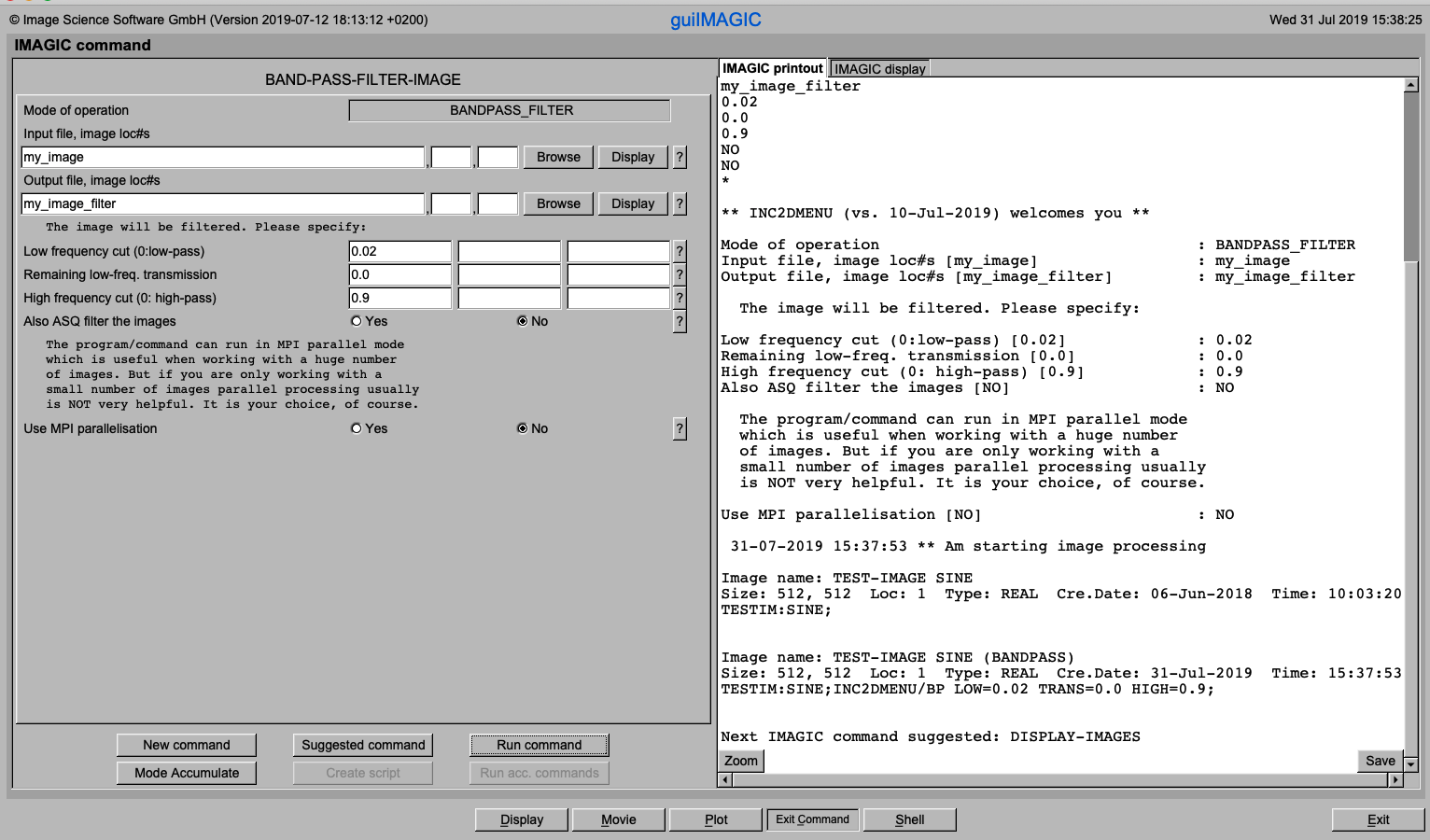
Start guiIMAGIC by giving guiIMAGIC in a terminal window. |
||||||||||||||||||||||||

|
GISP:
GISP guides you through a typical (ABC - alignment-by-classification) single particles image analysis in a GUI oriented environment. 
Start GISP by giving GISP in a terminal window. |

|
The IMAGIC image processing and analysis system: |
| Single Particles | |
| 3-D Processing and Reconstruction | |
| 2-D Crystals | |
|
Morphology
|
|

|
IMAGIC EM Package |

|
GISP: The GUI oriented IMAGIC Single Particles Analysis |

|
IMAGIC Shared Resources |

|
em2em: 3DEM Image Conversion Program |

|
FSC: Program to calculate the Fourier Shell Correlation
|

|
Manuals
|

|
References
|
Site Notice | Privacy Policy | webmaster(at)ImageScience.de